Published in: Engineering Applications of Artificial Intelligence, Vol. 172 (2026), Article 114291 DOI: 10.1016/j.engappai.2026.114291 Authors: Maria Bashir, Nasir Rahim, Shaker El-Sappagh, Omar Amin El-Serafy, Tamer Abuhmed Affiliations: Sungkyunkwan University · Gachon University · Galala University · Cairo University
Overview
Mild Cognitive Impairment (MCI) sits on the transitional continuum between healthy aging and Alzheimer’s Disease. Identifying which patients will progress (pMCI) versus remain stable (sMCI) is among the most clinically consequential and technically difficult problems in neuroimaging AI. MRI-DeepTrust addresses this challenge through a tissue-aware deep ensemble framework that independently models Gray Matter (GM), White Matter (WM), and Cerebrospinal Fluid (CSF) via Bayesian-optimized CNNs — while simultaneously and systematically evaluating three pillars of trustworthy AI: adversarial robustness, gender-based fairness, and visual explainability.
| Metric | Result |
|---|---|
| mAUC on ADNI (proposed ensemble) | 89% |
| Accuracy on ADNI (clean conditions) | 88% |
| Accuracy after adversarial attack (PGD) | 81% |
| Demographic Parity gap (proposed EL) | 0.01 |
| External validation on NACC (mAUC) | 81% |
Why Trustworthiness Matters for Alzheimer’s Diagnosis
Deep learning models for neuroimaging diagnosis have achieved remarkable accuracy on benchmark datasets. But accuracy alone is insufficient for clinical deployment. A diagnostic system that performs well on average may still fail for certain demographic subgroups, collapse under minor image perturbations, or provide predictions that clinicians cannot interpret or verify. In Alzheimer’s Disease prognosis, where diagnostic errors influence long-term treatment planning and patient counseling, these failures carry serious consequences.
Existing frameworks typically address one pillar of trustworthiness in isolation. MRI-DeepTrust fills this gap with a system designed from the ground up to be accurate, robust, fair, and explainable simultaneously — a clinician-in-the-loop workflow ensures that visual explainability outputs and model stability are qualitatively reviewed throughout.
“No prior study combines GM, WM, and CSF-specific CNNs optimized via Bayesian search with unified assessments of adversarial robustness, demographic fairness, and clinically guided visual interpretability in a single end-to-end diagnostic framework.”
Three Pillars
🛡 Robustness — PGD-based adversarial attack evaluation (ε=0.01, 10 steps) plus adversarial training at 20%, 35%, and 50% augmentation ratios. The ensemble reduces mAUC degradation to ~13% versus ~16–18% for single models.
⚖ Fairness — Gender-based evaluation using Demographic Parity (DP), Equal Opportunity (EO), and Group Sufficiency (GS). The proposed EL achieves DP=0.01, EO=0.02, GS=0.09 — the lowest disparities across all models.
🔍 Explainability — Dual-level: 3D brain surface renderings for global structural context, plus 2D Grad-CAM/MedCAM attention maps at the slice level — reviewed by a neurologist in a clinician-in-the-loop workflow.
The MRI-DeepTrust Framework
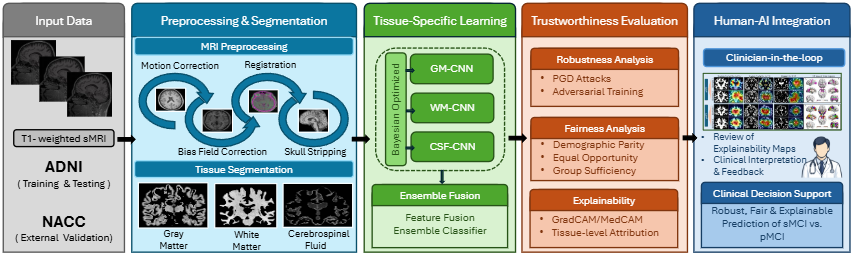
Figure 1. Block diagram of MRI-DeepTrust. MRI scans are preprocessed and segmented into GM, WM, and CSF. Separate Bayesian-optimized CNNs are trained per tissue, then fused through an ensemble classifier. Four dedicated modules evaluate trustworthiness: PGD robustness, adversarial training defense, gender fairness, and Grad-CAM explainability. External validation on NACC assesses cross-cohort generalizability.
MRI Preprocessing Pipeline
Each scan undergoes a standardized six-step pipeline before tissue segmentation:
-
Motion correction & reorientation — FSLEyes inspection and
fslreorient2stdcorrection - N4 bias field correction — ANTs N4 algorithm corrects intensity inhomogeneities
- MNI152 registration — Affine registration via FLIRT for spatial correspondence
- Skull stripping — BET2 from FSL with segmentation-based refinement
- Tissue segmentation — FSL 5.0 produces binary masks for GM, WM, and CSF
- Quality control — All masks visually inspected for anatomical accuracy
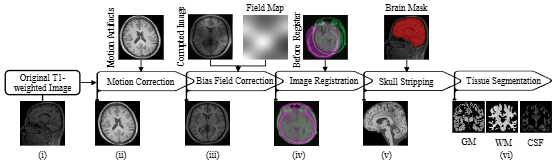
Figure 4. The complete MRI preprocessing pipeline from raw T1-weighted scan to tissue-segmented GM, WM, and CSF masks.
Bayesian-Optimized Tissue-Specific CNNs
A separate CNN is independently designed for each tissue type. Bayesian optimization searches over:
- Convolutional depth: 4–10 layers
- Filter counts: 96–320
- Kernel sizes: 3×3 or 5×5
- Dropout rates: 0.1–0.5
- Dense layer widths: 128–512
- Learning rates: 1e-2, 1e-3, 1e-4
This yields three architecturally distinct, complementary networks (see Table 1 in the paper for layer-wise configurations). After optimization, the final Softmax layers are removed, networks are frozen, and their representations are concatenated and passed through fully connected layers forming the ensemble classifier.
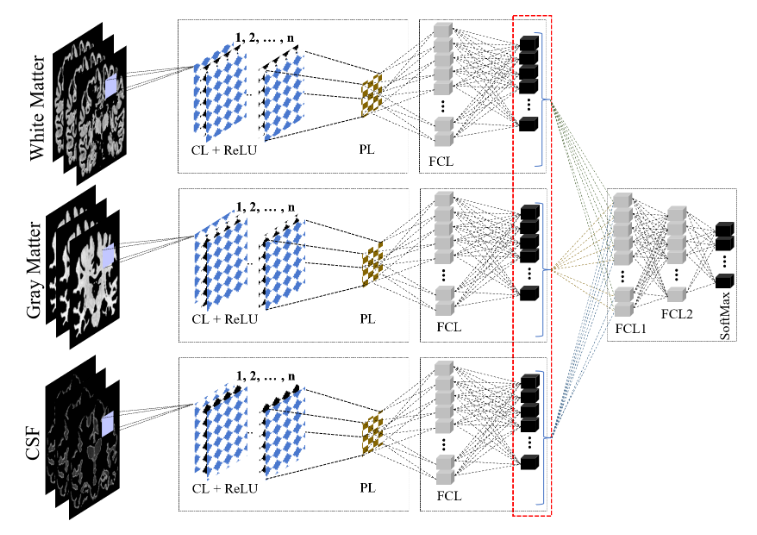
Figure 3. The proposed ensemble framework. Three tissue-specific CNNs (WM, GM, CSF) are trained independently, frozen, and fused through a fully connected ensemble classifier.
Eight Experiments — One Unified Evaluation
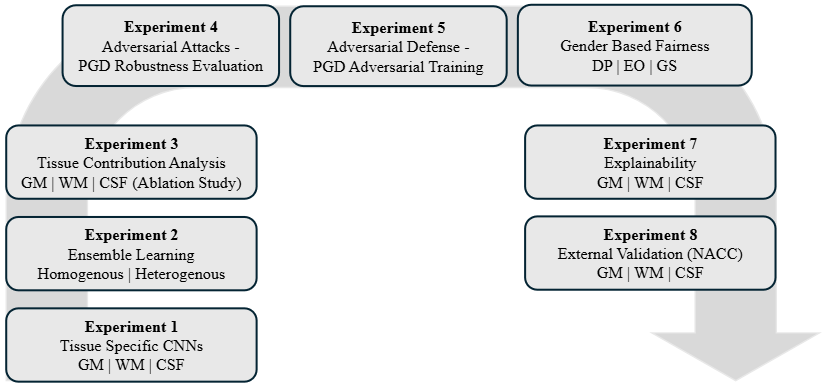
Figure 5. The structured eight-experiment evaluation pipeline.
| Experiment | Focus | Key Finding |
|---|---|---|
| Exp 1 | Tissue-specific CNNs | InceptionV3 best for GM; proposed model consistently competitive |
| Exp 2 | Ensemble learning | Proposed EL reaches 89% mAUC; heterogeneous > homogeneous |
| Exp 3 | Tissue ablation | GM removal: −20–25% mAUC; CSF removal: −5–12% mAUC |
| Exp 4 | PGD attack | Ensemble degrades ~13% vs. ~16–18% for base models |
| Exp 5 | Adversarial training | Proposed EL: 84% mAUC at 50% augmentation ratio |
| Exp 6 | Gender fairness | DP=0.01, EO=0.02, GS=0.09 (lowest across all models) |
| Exp 7 | Explainability | pMCI: focal, anatomically coherent activations; sMCI: diffuse |
| Exp 8 | NACC external validation | 81% mAUC without fine-tuning; no catastrophic degradation |
Results
Tissue-Specific and Ensemble Performance
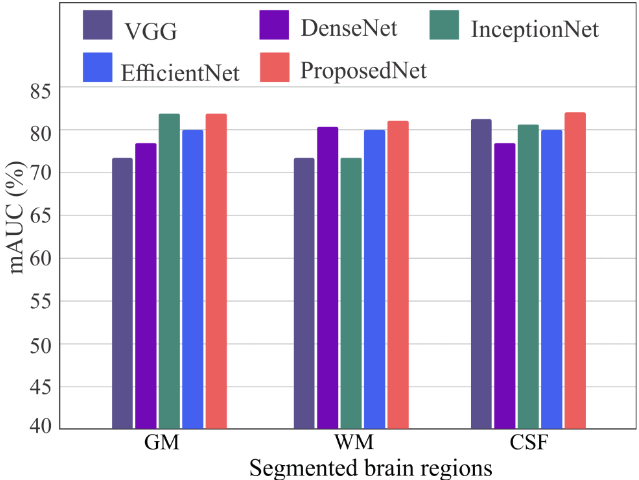
Figure 6. mAUC scores per tissue (GM, WM, CSF) across CNN architectures. EfficientNet and the proposed model maintain mAUC above 80% across all tissues, while other architectures exhibit tissue-dependent variability.
Ensemble comparison (ADNI, Accuracy):
| Model | Accuracy | F1 | AUC |
|---|---|---|---|
| Proposed EL (Heterogeneous) | 88% | 90% | 89% |
| Inception + Efficient + DenseNet | 89% | 88% | 87% |
| Ensemble DenseNet-121 | 86% | 86% | 85% |
| Ensemble EfficientNet | 81% | 84% | 84% |
| Ensemble VGG-16 | 78% | 81% | 82% |
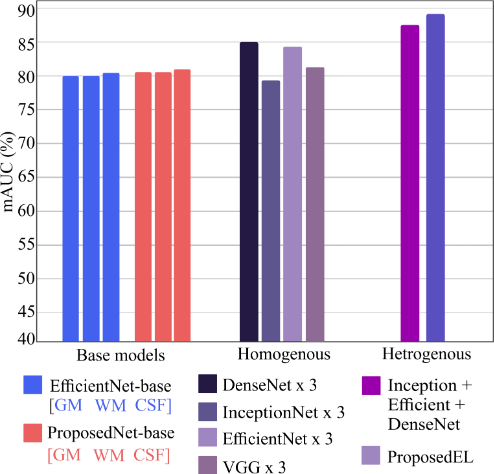
Figure 7. mAUC across base models, homogeneous, and heterogeneous ensembles. Heterogeneous configurations consistently provide the largest gains.
Tissue Ablation
All three tissues contribute meaningfully, but their roles differ:
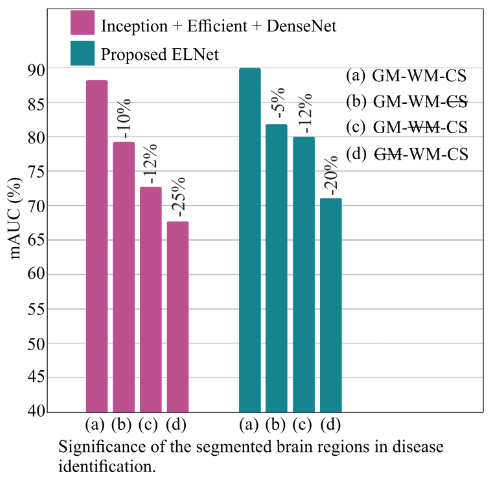
Figure 8. mAUC degradation when individual tissues are excluded. GM removal causes the largest drop (−20 to −25%), confirming its dominant role in discriminating pMCI from sMCI.
- GM removed: −20–25% mAUC — dominant contribution to classification
- WM removed: moderate degradation — deep and periventricular structural patterns
- CSF removed: −5–12% mAUC — ventricular and sulcal patterns still informative
Adversarial Robustness
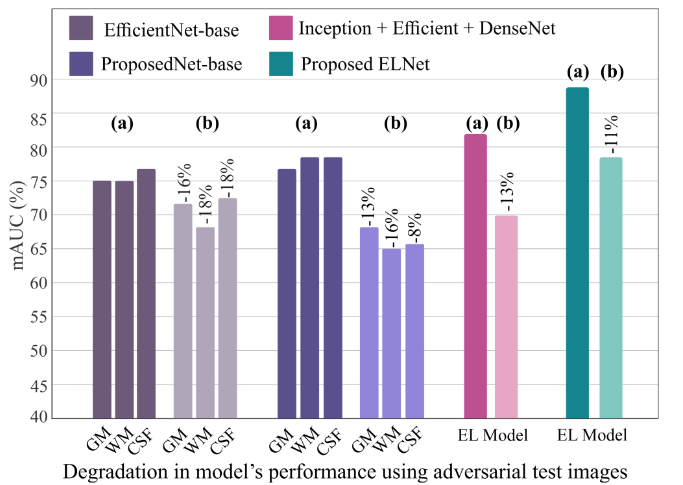
Figure 9. mAUC degradation under PGD attacks. The proposed EL (rightmost) exhibits the smallest drop (~11%), versus ~16–18% for base architectures and ~13% for the heterogeneous ensemble.
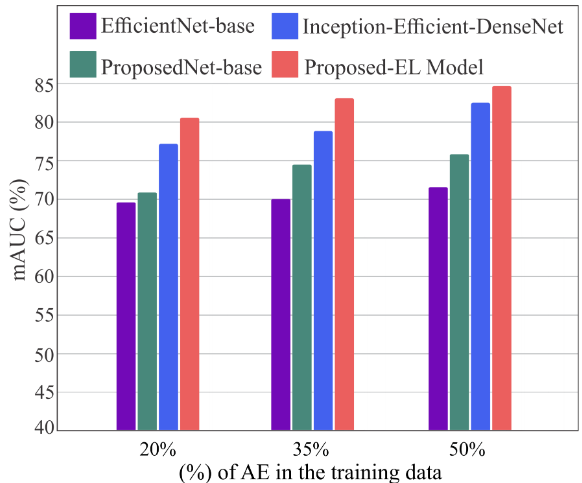
Figure 10. mAUC under adversarial training at 20%, 35%, and 50% augmentation. The proposed EL improves monotonically and reaches 84% mAUC at 50% augmentation — the only model that does not plateau.
Adversarial training at 50% augmentation:
| Model | Accuracy | mAUC |
|---|---|---|
| Proposed EL | 85% | 84% |
| Inception + Efficient + DenseNet | 83% | 82% |
| Proposed base (GM) | 78% | 77% |
| EfficientNet base (GM) | 75% | 73% |
Gender-Based Fairness
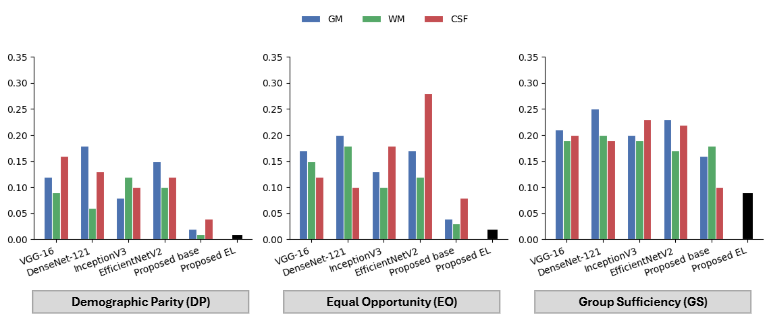
Figure 11. Demographic Parity (DP), Equal Opportunity (EO), and Group Sufficiency (GS) across all models. Lower values indicate smaller gender disparities. The proposed EL achieves the lowest values across all three metrics.
| Model | DP | EO | GS |
|---|---|---|---|
| Proposed EL | 0.01 | 0.02 | 0.09 |
| Proposed base (GM) | 0.02 | 0.04 | 0.16 |
| InceptionV3 (GM) | 0.08 | 0.13 | 0.20 |
| EfficientNetV2 (WM) | 0.10 | 0.12 | 0.17 |
| VGG-16 (GM) | 0.12 | 0.17 | 0.21 |
| DenseNet-121 (GM) | 0.18 | 0.20 | 0.25 |
Since all models were trained on identical data, fairness improvements stem from architectural design (tissue-specific learning + ensemble fusion) rather than dataset composition.
Clinician-Reviewed Explainability
MRI-DeepTrust employs a dual-level explainability strategy:
Global level — 3D surface rendering visualizes brain tissues across GM, WM, and CSF, offering anatomically meaningful views of cortical and subcortical regions commonly affected during disease progression.
Local level — 2D Grad-CAM/MedCAM attention maps highlight voxels within individual MRI slices that most strongly influence each prediction, overlaid directly on the original image.
Both outputs were reviewed by a neurologist in a clinician-in-the-loop workflow, confirming anatomical consistency and supporting translational feasibility.
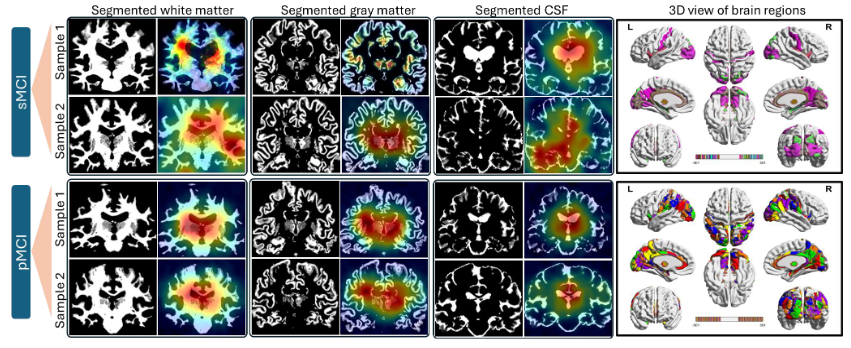
Figure 12. Tissue-specific Grad-CAM explainability maps for sMCI (top rows) and pMCI (bottom rows). sMCI attention patterns are diffuse and spatially distributed. pMCI patterns are stronger, more focal, and anatomically structured — involving the hippocampus, amygdala, fusiform gyrus, superior/middle temporal gyri, and periventricular regions. 3D surface renderings (rightmost) confirm the progressive shift from uncertain to decisive network attention as cognitive impairment advances.
External Validation on NACC
The ADNI-trained framework is directly evaluated on the independent NACC cohort — without fine-tuning — alongside ViT and CNN baselines. NACC presents greater scanner variability, demographic diversity, and labeling heterogeneity than ADNI.
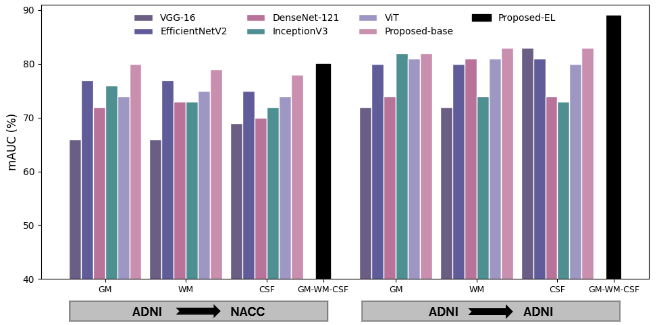
Figure 13. mAUC under internal (ADNI→ADNI) and external (ADNI→NACC) evaluation. The proposed framework consistently outperforms competing models in both settings.
| Model (Tissue) | ADNI mAUC | NACC mAUC | Drop |
|---|---|---|---|
| Proposed EL | 89% | 81% | −8% |
| EfficientNetV2 (GM) | 80% | 77% | −3% |
| InceptionV3 (GM) | 82% | 76% | −6% |
| ViT (GM) | 81% | 74% | −7% |
| VGG-16 (GM) | 72% | 66% | −6% |
Performance decline is consistent with expected domain shift — no catastrophic degradation — reinforcing the framework’s translational potential.
State-of-the-Art Comparison
MRI-DeepTrust is the only framework in the field to simultaneously evaluate all three trustworthiness pillars (Exp = Explainability, Fair = Fairness, Rob = Robustness).
| Study | Method | Accuracy | Exp | Fair | Rob |
|---|---|---|---|---|---|
| MRI-DeepTrust (Ours) | MRI-DeepTrust | 88% / 81%* | ✅ | ✅ | ✅ |
| Leonardsen et al. 2024 | CNN + LRP | 84% | ✅ | — | — |
| Vlontzou et al. 2025 | SVM | 87% | ✅ | — | — |
| Tong et al. 2025 | ML-Models | 76% | ✅ | ✅ | — |
| Zhang et al. 2023 | DAUF | 76% | ✅ | — | — |
| Ahmed et al. 2019 | EL-Model | 86% | — | — | — |
*88% before adversarial attack / 81% after adversarial attack (ε=0.01, PGD).
Conclusion
MRI-DeepTrust demonstrates that it is possible — and necessary — to design neuroimaging AI systems that are simultaneously accurate, robust, fair, and explainable. By independently modeling GM, WM, and CSF through Bayesian-optimized CNNs and fusing their representations through a heterogeneous ensemble, the framework captures complementary tissue-specific disease signatures while naturally reducing demographic disparities and improving resilience to adversarial perturbations.
The clinician-in-the-loop validation confirms anatomical consistency of the model’s attention patterns, bridging the gap between algorithmic performance and clinical relevance. External validation on NACC demonstrates stable cross-cohort generalization — a prerequisite for real-world deployment.
Future directions: extending fairness to ethnicity and socioeconomic attributes; stronger adversarial defenses; multimodal fusion with longitudinal and genetic data; validation across larger multi-center cohorts.
Citation
@article{mrideeptrust2026,
title = {Trustworthy Alzheimer's diagnosis: Integrating robustness, fairness,
and explainability in neuroimaging based deep ensemble framework},
author = {Bashir, Maria and Rahim, Nasir and El-Sappagh, Shaker and
El-Serafy, Omar Amin and Abuhmed, Tamer},
journal = {Engineering Applications of Artificial Intelligence},
volume = {172},
pages = {114291},
year = {2026},
doi = {10.1016/j.engappai.2026.114291}
}
Supported by the National Research Foundation of Korea (NRF) grant RS-2021-NR058558, IITP-ICT Creative Consilience Program (IITP-2025-RS-2020-II201821), and AI Platform grant No. 2022-0-00688.