Accepted at ICPR 2026 Authors: Firuz Juraev, Abdenour Soubih, Tamer Abuhmed · Sungkyunkwan University Code: github.com/InfoLab-SKKU/mm-des
Overview
Multimodal learning is crucial in medical applications, where patient data spans heterogeneous sources — skin images, physiological signals, radiology reports, and clinical notes. While multimodal ensemble models exploit complementary information, they rely on static model sets and assume all modalities are available at inference time. MM-DES is the first multimodal dynamic ensemble selection framework that identifies the most competent subset of models for each individual test sample, using contrastive multimodal representations to construct a sample-wise region of competence.
| Metric | Result |
|---|---|
| Accuracy on PAD-UFES-20 | 85.22% (vs. 83.26% best baseline) |
| AUROC on MIMIC-IV (all modalities) | 86.14% |
| AUROC gain over AutoPrognosis-M (MIMIC-IV) | +4.13% |
| Datasets evaluated | PAD-UFES-20, MIMIC-IV (Sepsis) |
| Modality types | Image · Tabular · Clinical Text |
The Problem with Static Multimodal Ensembles
In real clinical workflows, diagnostic decisions synthesize diverse data streams: a dermatologist reviews a smartphone photograph alongside patient history; an intensivist integrates radiology reports, vital-sign trends, and lab values. Multimodal machine learning mirrors this reality — but state-of-the-art approaches share a critical assumption: that the same set of models, with the same weights, is appropriate for every patient.
This assumption breaks down in practice. Modality quality varies across patients. Tabular features may be informative for one patient but noisy for another. An image-based classifier may be reliable for certain lesion types yet unreliable for others. And in the real world, modalities are frequently missing or corrupted — a patient may arrive without imaging, or a sensor may fail.
“Static ensembles apply the same models to every patient — regardless of which modalities are reliable for this individual.”
Dynamic Ensemble Selection (DES) addresses this by selecting and weighting classifiers at inference time, based on their performance on similar training samples. But DES has been studied almost exclusively for unimodal, tabular settings. MM-DES is the first framework to bring DES into the multimodal clinical domain.
The MM-DES Framework
MM-DES operates in two stages: a multimodal contrastive pretraining stage to build a shared embedding space, and a dynamic ensemble selection stage that performs sample-adaptive inference using modality-aware Regions of Competence.
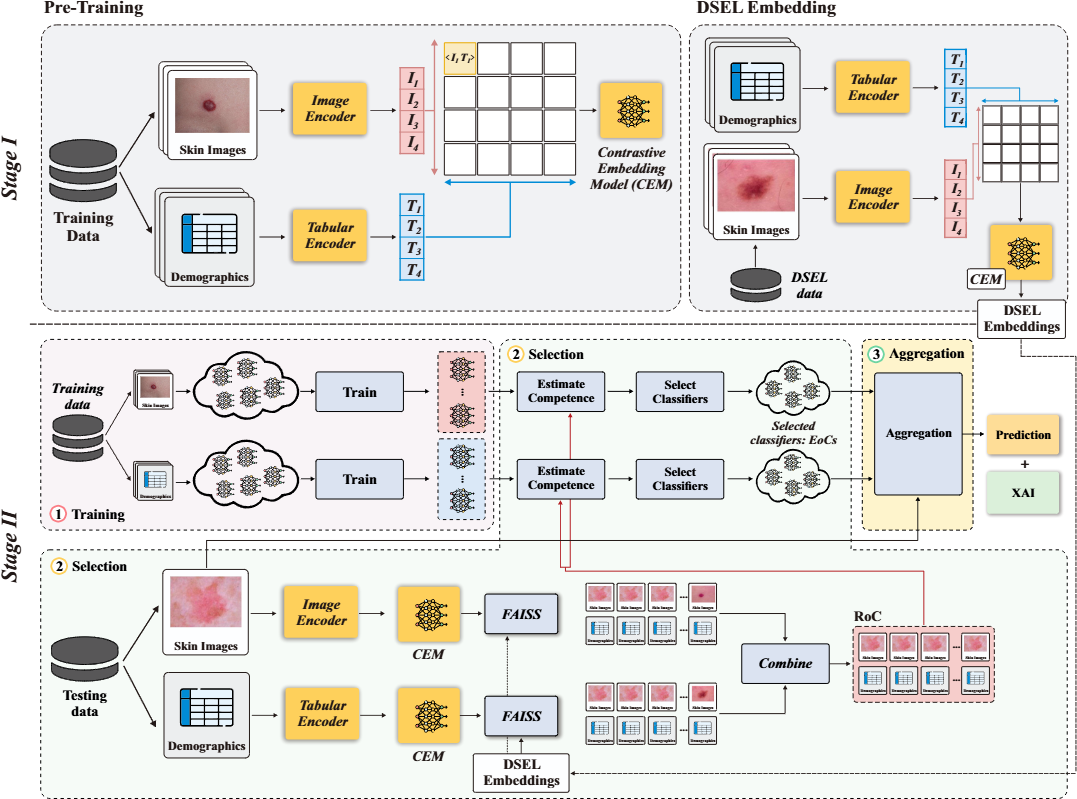
Figure 1. The MM-DES framework. Stage I (top): modality-specific encoders are jointly pretrained with the Symile contrastive objective to produce aligned multimodal embeddings, stored in a FAISS-indexed DSEL memory. Stage II (bottom): at inference time, modality-wise nearest-neighbor retrieval constructs a Region of Competence, used to estimate classifier competence and produce a competence-weighted multimodal prediction.
Stage I — Multimodal Contrastive Pretraining
Modality-specific encoders (image CNN, tabular MLP, language transformer) are jointly trained using the Symile contrastive objective. Unlike CLIP-style pairwise alignment, Symile generalizes contrastive learning to arbitrarily many modalities via multilinear inner products, treating each modality symmetrically as an anchor:
L_Symile = (1/M) Σ_m (1/B) Σ_i CE(ℓ_i^(m), i)
After convergence, the pretrained model is frozen and used to embed a Dynamic Selection Dataset (DSEL), which serves as the reference memory for downstream competence estimation.
Why Symile over CLIP? CLIP is architecturally limited to two modalities, making it unsuitable for MIMIC-IV’s three-modality setting. Ablation confirms Symile’s advantage across all metrics on PAD-UFES-20 (AUROC 96.99% vs 96.94%).
Stage II — Multimodal Dynamic Ensemble Selection
For each test patient:
- Embedding: the patient’s available modalities are encoded via the pretrained CEM, producing L2-normalized per-modality query embeddings.
- RoC Construction: independent FAISS searches in each modality’s embedding space retrieve the top-K most similar DSEL patients, with similarity scores aggregated across modalities using modality-specific weights.
- Competence Estimation: each classifier is scored by its local accuracy within the RoC — how often it correctly predicted the RoC patients’ labels.
-
Weighted Fusion: predictions are combined via competence- and modality-weighted aggregation:
p_q += w_m · κ_j^(m) · c_j^(m)(x_q^(m))
Missing modality handling: When a modality is absent, MM-DES zeros out its weight and constructs the RoC from available modalities only — no imputation, no retraining. This makes the framework naturally robust to partial and complete modality absence.
Datasets and Base Classifiers
PAD-UFES-20 — Skin Lesion Classification
A publicly available dataset of 2,298 smartphone-acquired skin lesion images paired with patient metadata across six lesion categories (BCC, MEL, SCC, ACK, NEV, SEK), with all cancer cases biopsy-confirmed.
Classifier pool (10 models):
- Image: ResNet-18, ResNet-34, EfficientNet-B0, MobileNet-V2, RegNet
- Tabular: Random Forest, CatBoost, XGBoost, MLP, SVC
Best single models: CatBoost (82.17% acc, 95.86% AUROC) · ResNet-18 (76.96% acc, 92.60% AUROC)
MIMIC-IV — Sepsis Prediction
A balanced cohort of 8,156 patients (4,078 sepsis / 4,078 non-sepsis) from the MIMIC-IV clinical database, combining three modalities: radiology report text (INDICATION section), structured vital-sign summaries, and chest X-ray images.
Classifier pool (11 models):
- Text: ClinicalBERT, PubMedBERT, BioBERT
- Tabular: CatBoost, XGBoost, Random Forest, MLP, SVC
- Image: ViT-Tiny, ResNet-18, MobileNet-V2
Results
PAD-UFES-20
MM-DES achieves the best results across all four metrics:
| Model | Modality | Accuracy | F1 | Bal Acc | AUROC |
|---|---|---|---|---|---|
| MM-DES (Ours) | MM | 85.22% | 79.73% | 76.20% | 96.99% |
| AutoPrognosis-M | MM | 83.26% | 77.65% | 74.51% | 96.70% |
| Static Ensemble (Soft) | Tabular | 82.39% | 71.82% | 66.91% | 96.07% |
| Best Tab (CatBoost) | Tabular | 82.17% | 74.40% | 70.16% | 95.86% |
| Best Img (ResNet-18) | Image | 76.96% | 70.42% | 66.13% | 92.60% |
The gain in balanced accuracy (+1.69% over AutoPrognosis-M) is particularly meaningful: dynamic, locally-estimated classifier selection helps overcome class imbalance that static ensembles struggle with.
MIMIC-IV Sepsis Prediction
| Model | Modality | Accuracy | F1 | Bal Acc | AUROC |
|---|---|---|---|---|---|
| MM-DES (Ours) | MM-LTI | 77.17% | 76.99% | 77.18% | 86.14% |
| MM-DES (Ours) | MM-LT | 75.98% | 75.78% | 75.98% | 83.86% |
| AutoPrognosis-M | MM-LTI | 74.34% | 74.11% | 74.32% | 82.01% |
| AutoPrognosis-M | MM-LT | 73.68% | 73.40% | 73.68% | 80.42% |
| Static Ensemble (Soft) | Text | 72.84% | 73.92% | 72.84% | 80.13% |
| Best Tab (CatBoost) | Tabular | 71.65% | 71.56% | 71.65% | 77.72% |
MM-LT = language + tabular; MM-LTI = language + tabular + image.
Robustness to Missing Modalities
A defining challenge in clinical deployment is modality absence. MM-DES handles this gracefully — by constructing the RoC from whatever modalities are available.
PAD-UFES-20
| Setting | Missing | Accuracy | F1 | Bal Acc | AUROC |
|---|---|---|---|---|---|
| Full | None | 85.22% | 79.73% | 76.20% | 96.99% |
| Image only | Tabular (100%) | 77.61% | 70.40% | 67.81% | 93.20% |
| Tabular only | Image (100%) | 82.83% | 72.65% | 67.71% | 96.20% |
| Partial | Tabular (20%) | 82.10% | 76.20% | 72.90% | 95.40% |
| Partial | Image (20%) | 84.10% | 78.10% | 74.80% | 96.50% |
Under a realistic 20% partial-missing rate, performance remains within 1–3% of the fully-observed baseline. The system does not over-rely on any single modality.
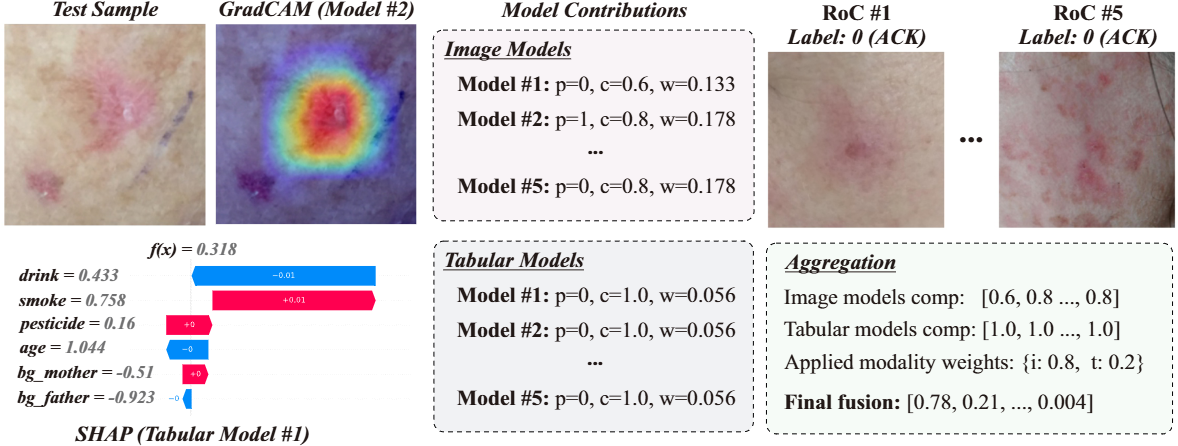

Instance-Level Interpretability
MM-DES provides multi-level interpretability for every prediction:
Model contributions: for each classifier in the pool, MM-DES reports the prediction, competence score κ, and assigned weight w — revealing which models the ensemble relied on and why.
Modality-level insights: the applied modality weights are reported explicitly, showing whether the final decision was driven by imaging, tabular variables, or clinical text.
Feature-level explanations:
- Image models → Grad-CAM highlights salient lesion regions
- Tabular models → SHAP quantifies feature contributions (e.g., smoking history, age, vital signs)
- Language models → CAPTUM attributes importance to specific tokens in clinical notes
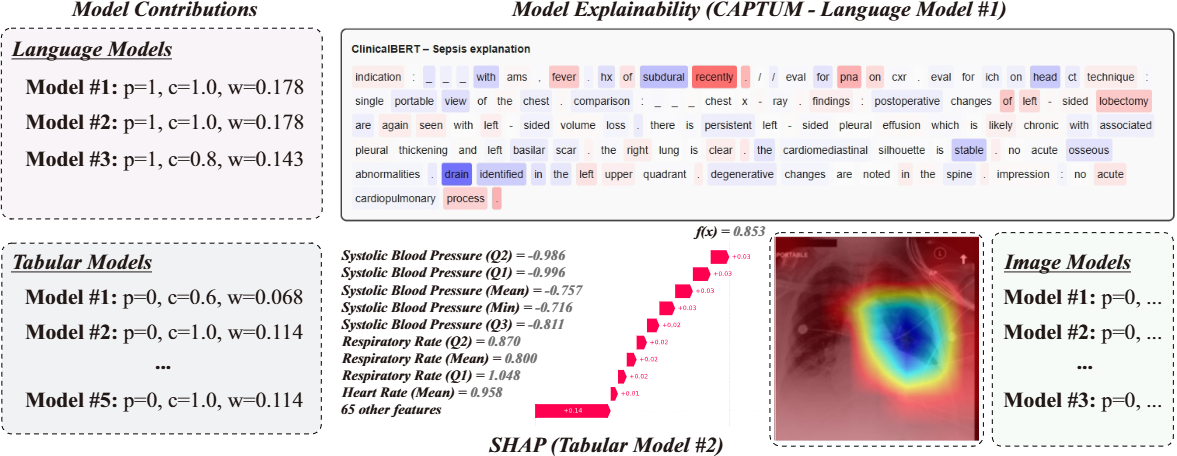
Figure 3. Interpretability output for a PAD-UFES-20 test sample. Despite image models initially receiving higher weights, competence analysis reveals that tabular models perform better locally and correctly drive the prediction. SHAP shows smoking history and age are the most important features; Grad-CAM highlights the lesion region.
Ablation Highlights
Neighborhood size (k): Performance peaks at k=10 and plateaus. Smaller neighborhoods (k=5) introduce instability; larger ones (k=15) dilute locality. Default: k=10.
| k | Accuracy | F1 | Bal Acc | AUROC |
|---|---|---|---|---|
| 5 | 84.78% | 79.30% | 75.52% | 96.43% |
| 7 | 85.00% | 79.24% | 75.55% | 96.64% |
| 10 | 85.22% | 79.73% | 76.20% | 96.99% |
| 12 | 85.03% | 79.63% | 76.11% | 96.94% |
| 15 | 85.03% | 79.63% | 76.11% | 96.94% |
Modality weights: Emphasizing tabular slightly more (0.4 image, 0.6 tabular) yields the best results for PAD-UFES-20, reflecting the higher discriminative power of structured clinical variables.
| Img | Tab | Accuracy | AUROC |
|---|---|---|---|
| 0.2 | 0.8 | 84.35% | 94.51% |
| 0.3 | 0.7 | 84.78% | 95.02% |
| 0.4 | 0.6 | 85.22% | 96.99% |
| 0.5 | 0.5 | 84.13% | 95.91% |
| 0.6 | 0.4 | 84.04% | 94.98% |
Search backend: FAISS (67 ms, 1.8 MB) outperforms both HNSW (70 ms, 2.1 MB) and brute-force K-NN (970 ms, 1.8 MB) on latency — the clear choice for clinical deployment.
Conclusion
MM-DES demonstrates that Dynamic Ensemble Selection, previously confined to unimodal tabular settings, can be successfully extended to multimodal clinical prediction. By grounding the Region of Competence in contrastive multimodal embeddings, MM-DES achieves sample-adaptive model selection that naturally handles missing modalities, improves over static multimodal ensembles, and remains interpretable at the instance level.
The results across PAD-UFES-20 and MIMIC-IV show consistent improvements over AutoPrognosis-M and unimodal baselines, with the largest gains in challenging missing-modality and class-imbalanced scenarios — precisely the conditions most common in real clinical deployment.
Future directions: adaptive per-sample modality weighting, uncertainty-aware DES parameter selection, richer modality-contribution analyses, and extension to larger clinical datasets.
Citation
@inproceedings{mmdes2026,
title = {MM-DES: Enhancing Multimodal Clinical Prediction
with Joint Contrastive Embeddings and Dynamic Ensembles},
author = {Juraev, Firuz and Soubih, Abdenour and Abuhmed, Tamer},
booktitle = {Proceedings of the International Conference on
Pattern Recognition (ICPR)},
year = {2026}
}
This work was supported by the National Research Foundation of Korea (NRF) grant RS-2021-NR058558 and IITP-ICT Creative Consilience Program (IITP-2026-RS-2020-II201821).